Research
Overview
I develop signal processing and statistical modeling approaches to study brain–body dynamics during sleep and in clinical settings. My work focuses on multimodal electrophysiology (EEG + EGG + autonomic signals), wearable sensing, and interpretable machine learning, with the broader goal of building scalable biomarkers of physiology and health.
Brain-Body Coupling During Human Healthy Sleep
Question: How do peripheral rhythms coordinate with neural sleep dynamics, and what can this tell us about sleep physiology and cognition?
I analyze simultaneous high-density EEG and electrogastrography (EGG) recorded overnight to characterize stomach–brain coupling across sleep stages and timescales. This work tests whether gastric rhythms provide structured timing and infraslow organization that aligns with cortical events and sleep microarchitecture.
Methods: time-frequency analysis, phase-based coupling metrics, mixed-effects models, permutation testing
Keywords: sleep microarchitecture • interoception • infraslow dynamics • spindles and slow wave oscillations
Selected work: Simultaneous stomach–brain electrophysiology reveals dynamic coupling in human sleep
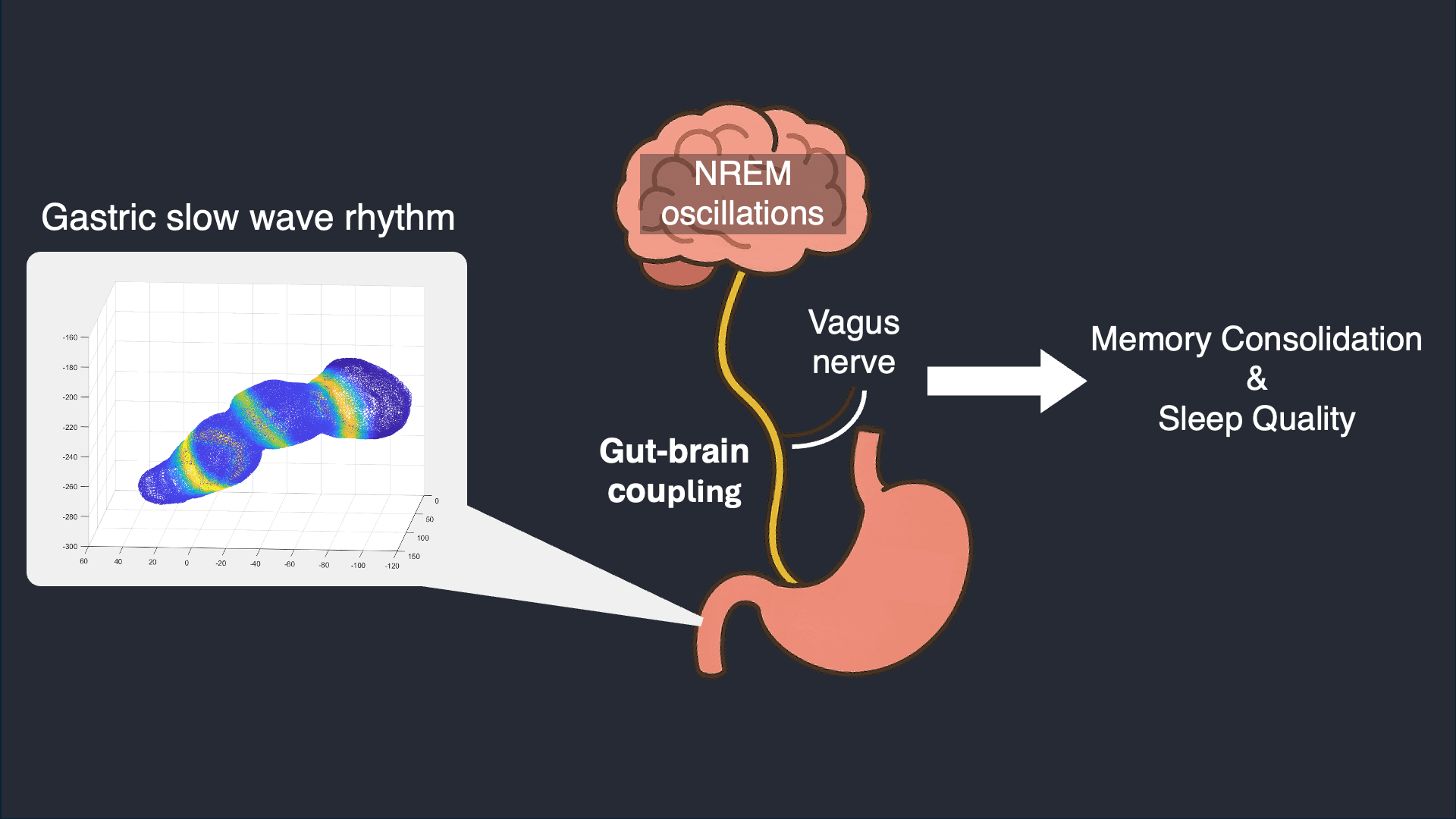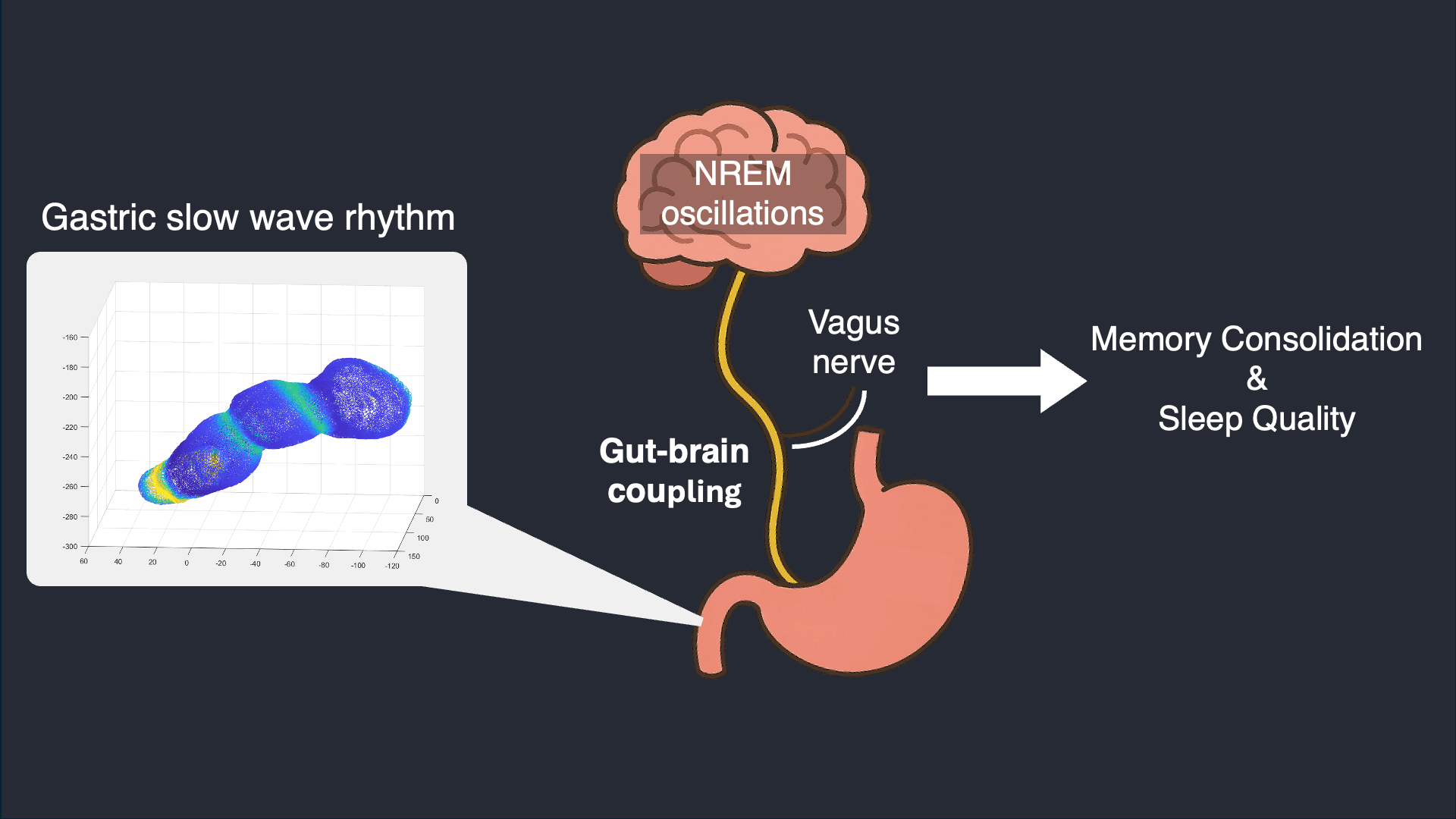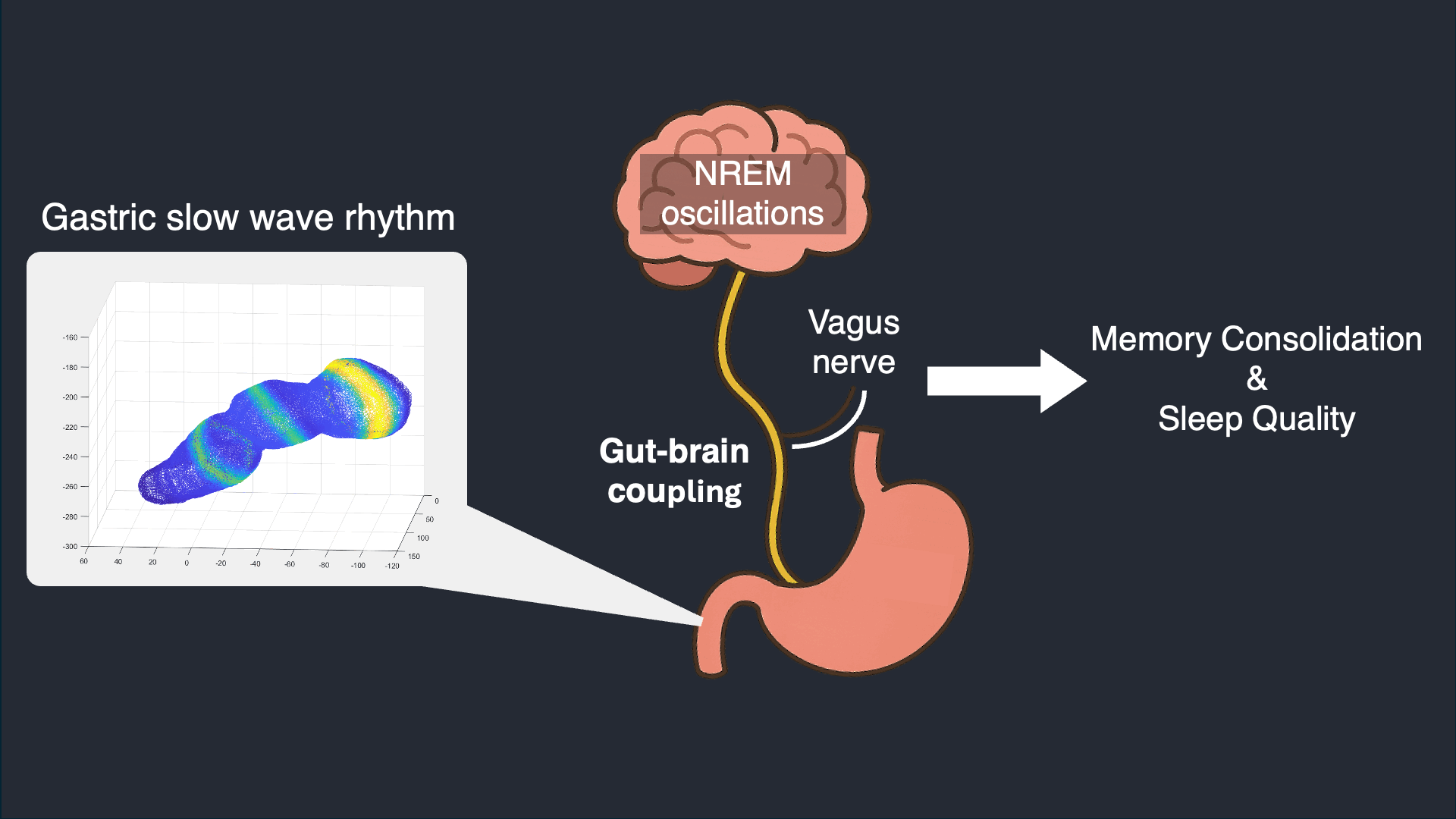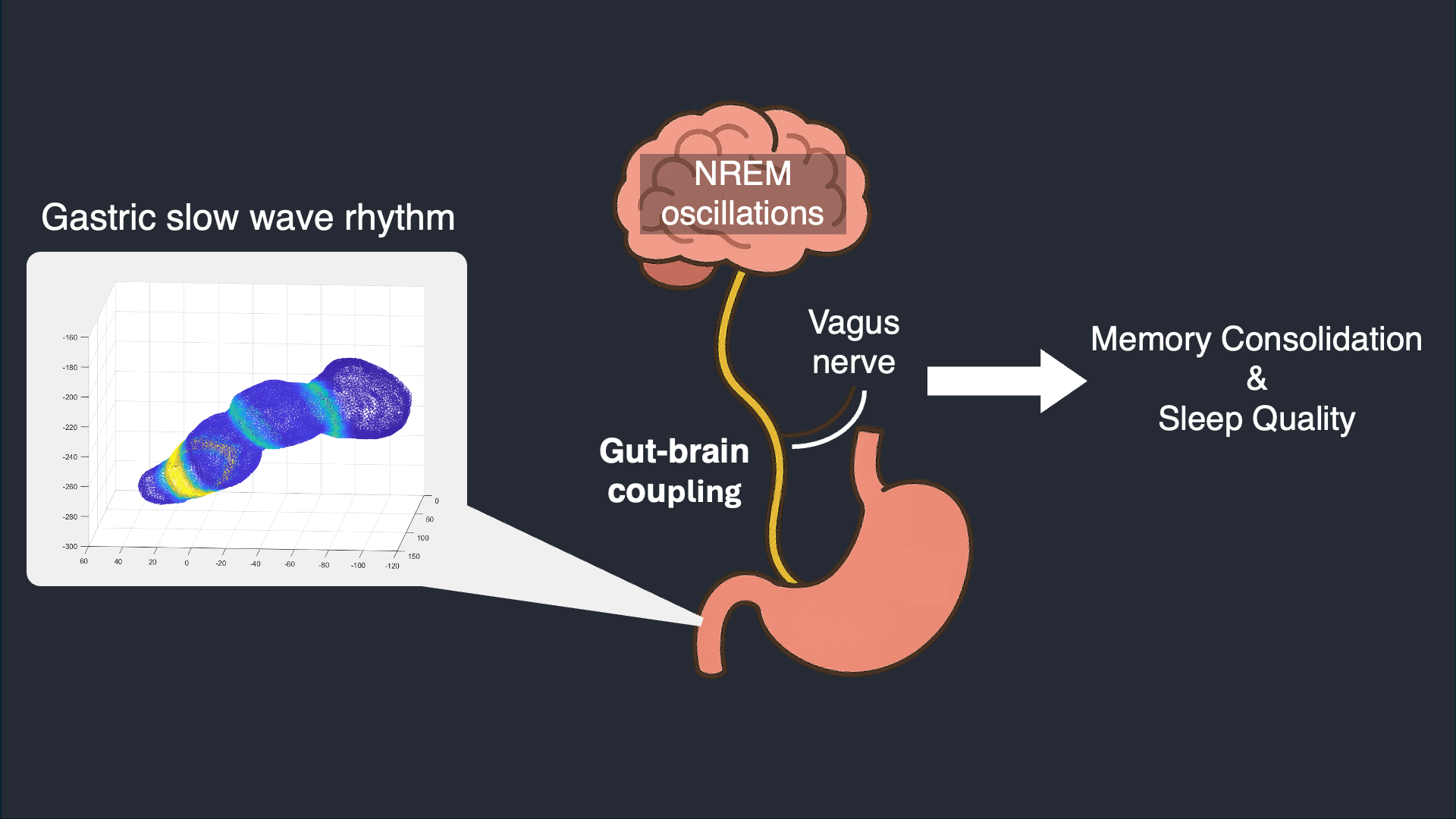
Wearable Autonomic Signal Processing
Question: How can we robustly quantify autonomic physiology from wearable sensors to support real-world monitoring?
I build pipelines for extracting interpretable autonomic features (e.g., HR/HRV dynamics) from wearable ECG/PPG, and evaluate how these signals vary across behavioral states and clinical contexts. I’m interested in building reliable metrics that generalize across devices and noisy environments.
Methods: signal quality indexing, robust feature extraction, validation & benchmarking
Keywords: HRV • wearable sensing • robustness • translational biomarkers
Interpretable Machine Learning for Facial Palsy
Question: Can computer vision and machine learning quantify dynamic facial function and predict clinical outcomes?
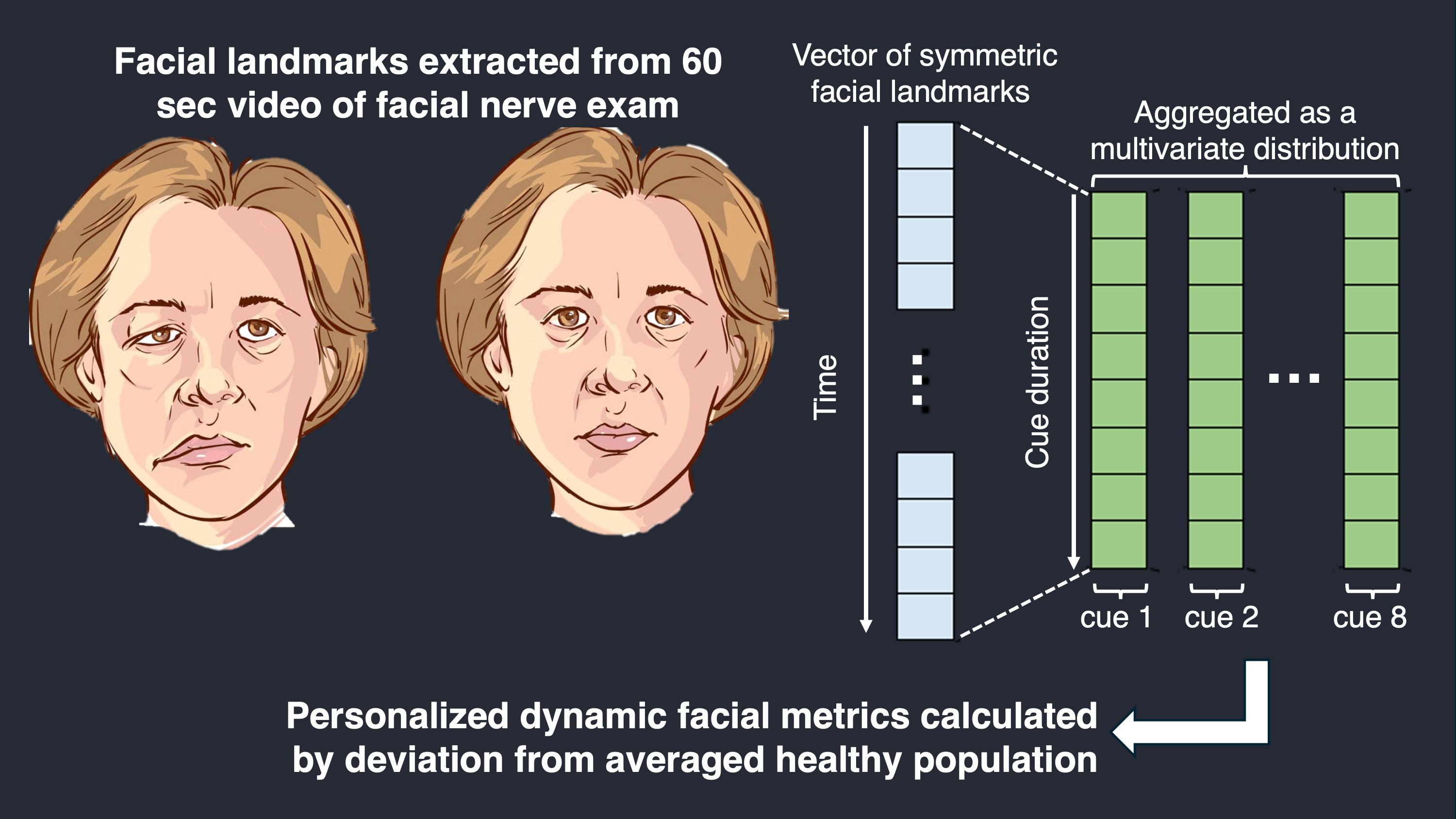
I developed machine learning methods employing likelihood ratio tests, optimal transport theory, and Mahalanobis distances to measure facial movement dynamics from video and to predict facial palsy outcomes using clinically meaningful targets. This work aims to support surgeon decision-making and longitudinal tracking.
Methods: landmark-based dynamics, ordinal regression, model evaluation/validation
Selected work: Machine Learning Methods to Track Dynamic Facial Function in Facial Palsy
Earlier Work: Bioelectronics for Cardiac Tissue Engineering
I previously worked on integrated bioelectronic platforms for monitoring and modulating engineered cardiac tissues, including heart-on-a-chip systems and optogenetic + bioelectronic interfaces for long-term stimulation and recording.
Collaborations & Tools
- Data processing in Python (MNE, NumPy/SciPy), statistical modeling (statsmodels, scikit-learn, PyTorch), reproducible pipelines (HPC + Git)
- Open to collaborations on multimodal physiology, wearable analytics, and clinical translation
